Home
About CBI
Now more than ever before, the power to turn huge volumes of data into information about our world has put the answers to some of life’s most challenging questions within our grasp. The George Washington University’s Computational Biology Institute (CBI) brings together leading faculty in biology, medicine and computing to harness this information, opening new doors of discovery that have the potential to benefit millions. CBI is also maximizing the university’s unique relationships in the nation’s capital to form research partnerships and spotlight cutting-‐edge topics that enhance human and environmental health. By blending its own skills with the expertise of engineers, mathematicians, statisticians, clinicians and others, the CBI is contributing to knowledge and resources that are used by researchers on a global scale, influencing how the world uses science and technology to solve its most pressing problems. With these truly incomparable resources and expertise, the CBI performs cutting-edge research and helps raise awareness of scientific advancements that improve our health, environment and overall quality of life.
RANKED #11
U.S. News & World Report's List of Best Public Health Graduate Programs
LOCATION
Washington D.C.'s only Public Health School
RESEARCH
Shaping Public Health Policy and Practice
FACULTY
130 Full-time Faculty Leading our Students, and 300 Part-time Faculty.
Featured Publications

Estimating rare disease prevalence and costs in the USA: a cohort study approach using the Healthcare Cost Institute claims data
April 26, 2024
Estimating rare disease prevalence and costs in the USA: a cohort study approach using the Healthcare Cost Institute claims data
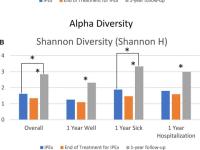
Impact of Antibiotics on the Lung Microbiome and Lung Function in Children With Cystic Fibrosis 1 Year After Hospitalization for an Initial Pulmonary Exacerbation
April 26, 2024
The purpose of this study was to evaluate changes in the microbiome and lung function in people with CF over 1 year after an initial pulmonary exacerbation.

Integrated Fertilization with Bagasse Vermicompost Changes the Microbiome of Mencía Must and Wine
April 26, 2024
Grape marc, or bagasse, is a byproduct of wine production that can be vermicomposted and used as organic fertilizer, which improve grape yields and wine quality

Nasopharyngeal metatranscriptome profiles of infants with bronchiolitis and risk of childhood asthma: a multicentre prospective study
April 26, 2024
Our objectives were to identify metatranscriptome profiles of infant bronchiolitis, and to examine relationship with host transcriptome and asthma development.