Home
About CBI
Now more than ever before, the power to turn huge volumes of data into information about our world has put the answers to some of life’s most challenging questions within our grasp. The George Washington University’s Computational Biology Institute (CBI) brings together leading faculty in biology, medicine and computing to harness this information, opening new doors of discovery that have the potential to benefit millions. CBI is also maximizing the university’s unique relationships in the nation’s capital to form research partnerships and spotlight cutting-‐edge topics that enhance human and environmental health. By blending its own skills with the expertise of engineers, mathematicians, statisticians, clinicians and others, the CBI is contributing to knowledge and resources that are used by researchers on a global scale, influencing how the world uses science and technology to solve its most pressing problems. With these truly incomparable resources and expertise, the CBI performs cutting-edge research and helps raise awareness of scientific advancements that improve our health, environment and overall quality of life.
RANKED #11
U.S. News & World Report's List of Best Public Health Graduate Programs
LOCATION
Washington D.C.'s only Public Health School
RESEARCH
Shaping Public Health Policy and Practice
FACULTY
130 Full-time Faculty Leading our Students, and 300 Part-time Faculty.
Featured Publications

Analysis of metagenomic data
August 19, 2025
Metagenomics has revolutionized our understanding of microbial communities, offering unprecedented insights into their genetic and functional diversity across Earth’s diverse ecosystems.

Locus-specific HERV expression associated with hepatocellular carcinoma
August 19, 2025
Human endogenous retroviruses (HERVs) harbor accessory proteins that influence cellular processes and have been linked to a wide variety of diseases, including cancer. This study investigates locus-specific HERV expression and its association with gene dysregulation in hepatocellular carcinoma (HCC), a highly prevalent and deadly form of liver cancer worldwide.

Estimating rare disease prevalence and costs in the USA: a cohort study approach using the Healthcare Cost Institute claims data
March 5, 2024
Estimating rare disease prevalence and costs in the USA: a cohort study approach using the Healthcare Cost Institute claims data
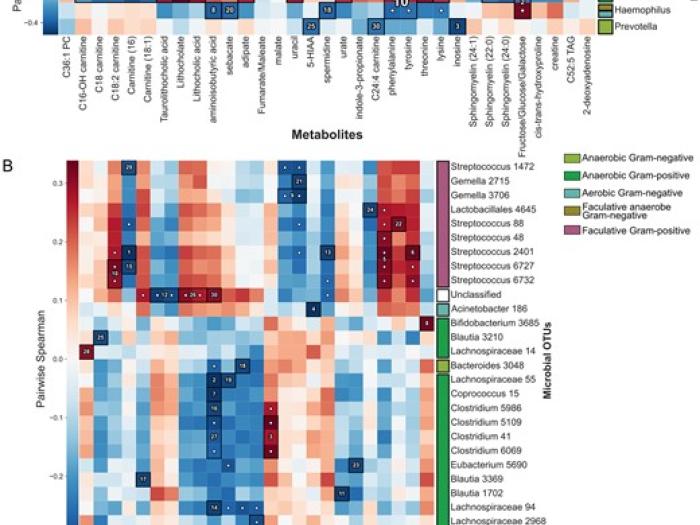
High-sensitivity pattern discovery in large, paired multiomic datasets
June 27, 2022
We present a novel framework which integrates hierarchical hypothesis testing with FDR correction to reveal linear and non-linear relationships among data.








